“A Bayesian approach to soil biogeochemical model evaluation [using information criteria and approximate cross-validation]”
This one took a while to get out. The project was started in late 2016 as an offshoot of Adriana Romero-Olivares' soil warming meta-analysis detailed in “Soil microbes and their response to experimental warming over time: A meta-analysis of field studies” and was originally meant to be a section of that paper before becoming its own project. I was slowed down in the interceding years of course by involvement in an ill-fated startup. The staccato and delayed research progression was of benefit in some ways due to the arrival of new updates and tools, such as the release of loo package.
I am glad to have finally gotten this paper out with the help of my advisor, Steve Allison, and the other authors, along with members of the Stan development team, such as Aki Vehtari, Michael Betancourt, Bob Carpenter, and Ben Bales. However, there is one thing I would like to take back and change, which is the title of the manuscript. I wish I had added the phrase “using information criteria and approximate cross-validation” at the end of the title to make it less vague.
Further summarizing and consolidating from the abstract, the general result of the paper was that a linear microbe-implicit soil biogeochemical system representing carbon (C) mass transfer in soil with first-order decay functions quantitatively outperformed a non-linear microbial-explicit system using Michaelis-Menten functions to represent C transfer in terms of widely applicable information criterion (WAIC) and leave-one-out cross-validation (LOOCV), conditional on the meta-analysis data set and prior distributions used.
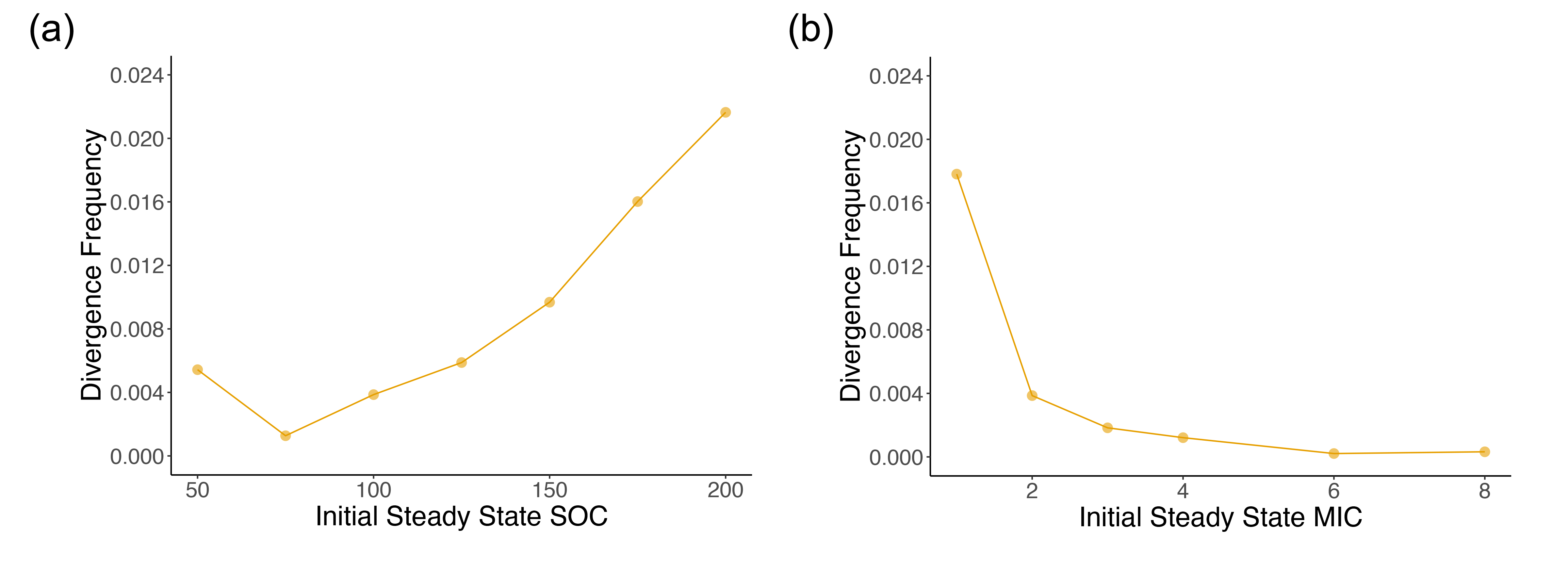
We did not explore this too deeply, but one observation that caught my eye that I would like to investigate more deeply in the future is the changing in the frequency of divergent transitions (which can be thought of as errors occurring during the Bayesian Hamiltonian Monte Carlo exploration due to constrained parameter space geometry) with the varying of initial conditions for the non-linear model. As can be seen in Figure S9 in the supplement, increasing the initial soil organic C in the model generally increased the frequency of divergences, whereas increasing the initial microbial biomass C decreased the the divergences.
As of now, I have not built an accompanying RShiny application for this manuscript due to the use of Stan, model instability for the non-linear system at some parameter regimes, and time constraints, but an RShiny application may follow for subsequent manuscripts in my thesis that build on this first chapter. Otherwise, the R and Stan code corresponding to this project is available for reference here via OSF or here via Github. For those looking for examples of implementing Earth system or biogeochemical models in Stan or use of loo for differential equation models, I hope the code can be of help.